A Short Introduction to splines2
Wenjie Wang
2025-02-28
Source:vignettes/splines2-intro.Rmd
splines2-intro.RmdIntroduction
The R package splines2 is intended to be a
user-friendly supplementary package to the base package
splines. It provides functions to construct a variety
of regression spline basis functions that are not available from
splines. Most functions have a very similar user
interface with the function splines::bs(). More
specifically, splines2 allows users to construct the
basis functions of
- B-splines
- M-splines
- I-splines
- C-splines
- periodic splines
- natural cubic splines
- generalized Bernstein polynomials
along with their integrals (except C-splines) and derivatives of given order by closed-form recursive formulas.
Compared to splines, the package
splines2 provides convenient interfaces for spline
derivatives with consistent handling on NA’s. Most of the
implementations are in C++ with the help of
Rcpp and RcppArmadillo since v0.3.0,
which boosted the computational performance.
In the remainder of this vignette, we illustrate the basic usage of most functions in the package through examples. We refer readers to Wang and Yan (2021) for a more formal introduction to the package with applications to shape-restricted regression. See the package manual for more details about function usage.
library(splines2)
packageVersion("splines2")## [1] '0.5.4'B-splines
B-spline Basis Functions
The bSpline() function generates the basis matrix for
B-splines and extends the function bs() of the package
splines by providing 1) the piece-wise constant basis
functions when degree = 0, 2) the derivatives of basis
functions for a positive derivs, 3) the integrals of basis
functions if integral = TRUE, 4) periodic basis functions
based on B-splines if periodic = TRUE.
One example of linear B-splines with three internal knots is as follows:
knots <- c(0.3, 0.5, 0.6)
x <- seq(0, 1, 0.01)
bsMat <- bSpline(x, knots = knots, degree = 1, intercept = TRUE)
plot(bsMat, mark_knots = "all")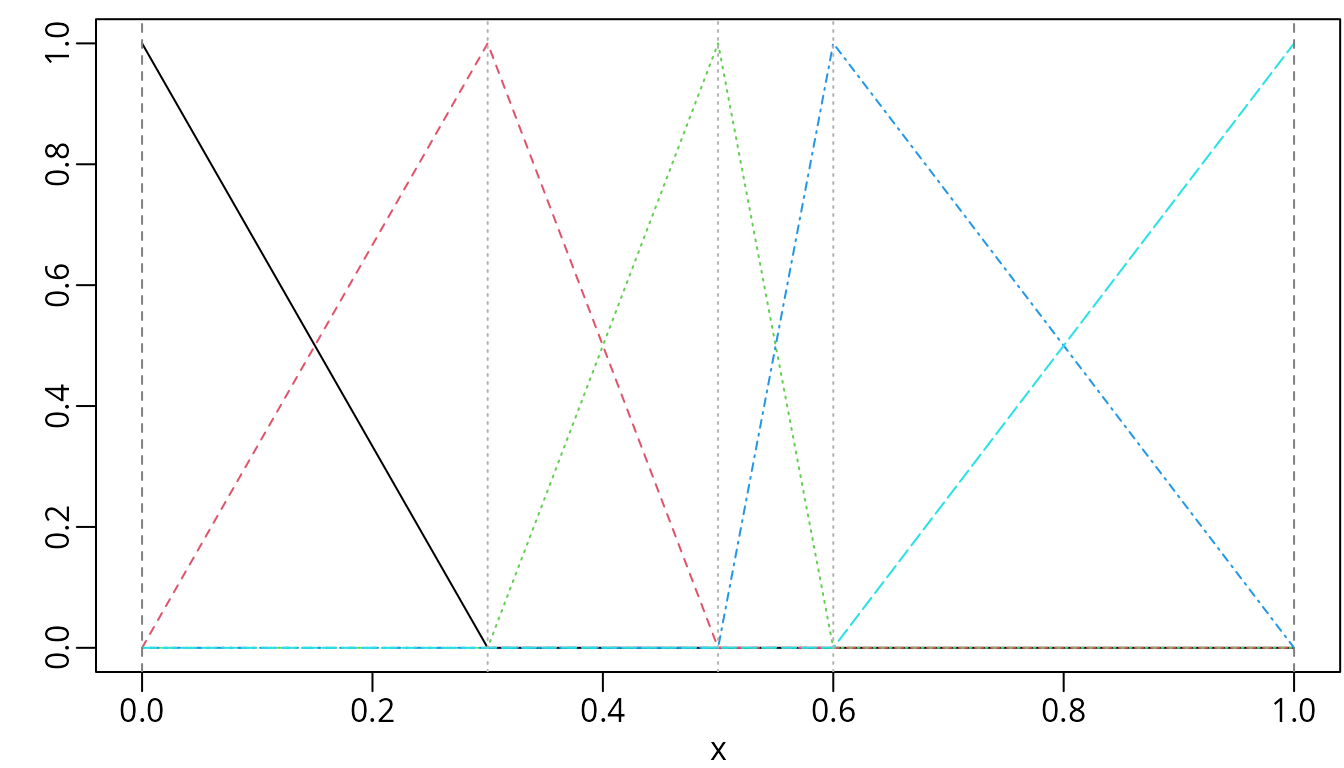
B-splines of degree one with three internal knots placed at 0.3, 0.5, and 0.6.
Integrals and Derivatives of B-splines
For convenience, the package also provides functions
ibs() and dbs() for constructing the B-spline
integrals and derivatives, respectively. Two toy examples are as
follows:
ibsMat <- ibs(x, knots = knots, degree = 1, intercept = TRUE)
op <- par(mfrow = c(1, 2))
plot(bsMat, mark_knots = "internal")
plot(ibsMat, mark_knots = "internal")
abline(h = c(0.15, 0.2, 0.25), lty = 2, col = "gray")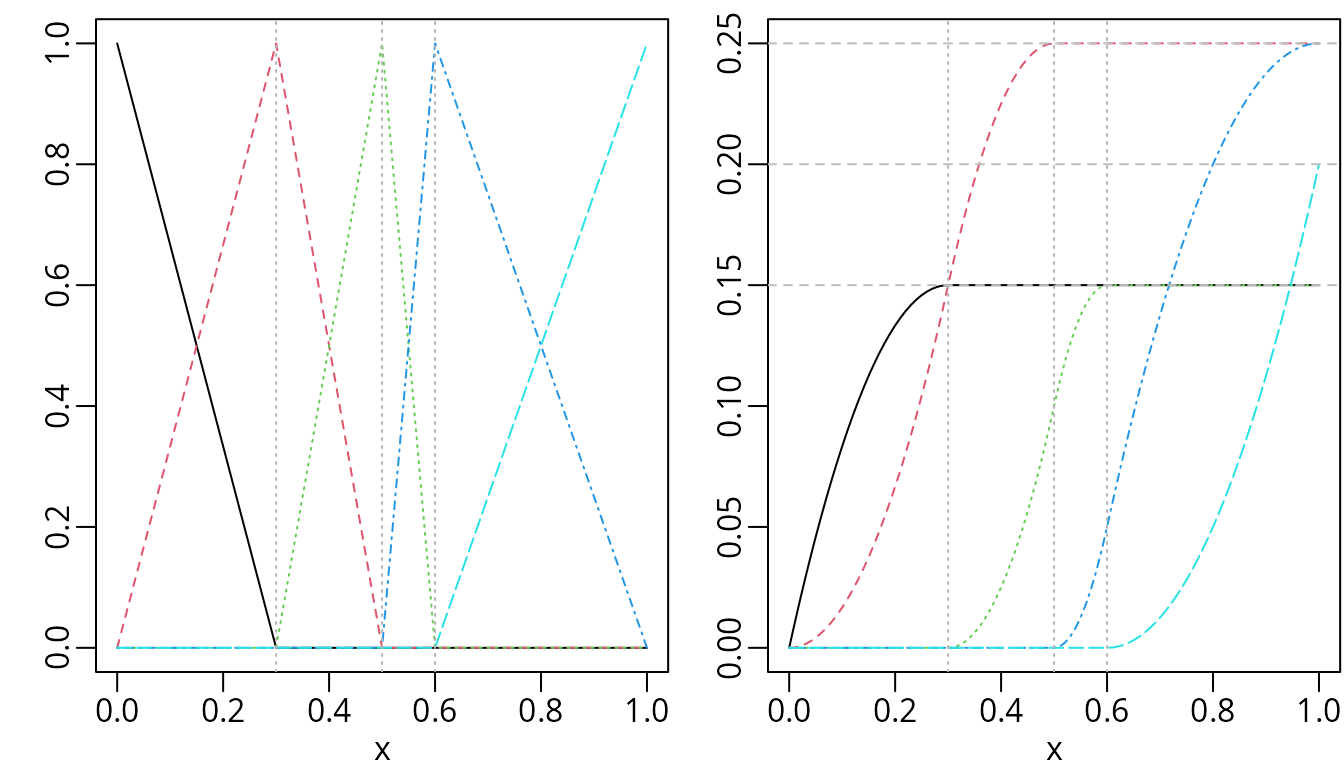
Piecewise linear B-splines (left) and their integrals (right).
bsMat <- bSpline(x, knots = knots, intercept = TRUE)
dbsMat <- dbs(x, knots = knots, intercept = TRUE)
plot(bsMat, mark_knots = "internal")
plot(dbsMat, mark_knots = "internal")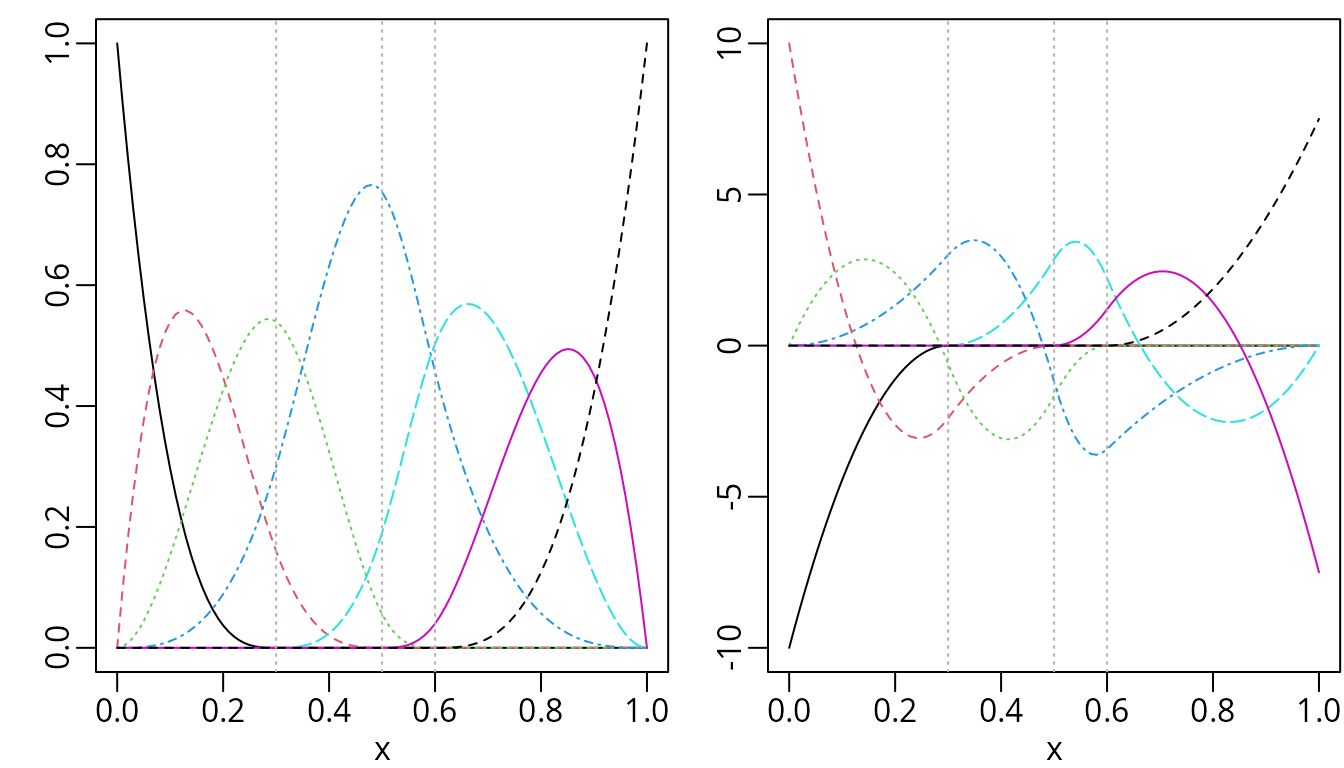
Cubic B-spline (left) and their first derivative (right).
We may also obtain the derivatives easily by the deriv()
method as follows:
Periodic B-splines
The function bSpline() produces periodic spline basis
functions following Piegl and Tiller (1997, chap.
12) when periodic = TRUE is specified. Different
from the regular basis functions, the x is allowed to be
placed outside the boundary and the Boundary.knots defines
the cyclic interval. For instance, one may obtain the periodic cubic
B-spline basis functions with cyclic interval (0, 1) as follows:
px <- seq(0, 3, 0.01)
pbsMat <- bSpline(px, knots = knots, Boundary.knots = c(0, 1),
intercept = TRUE, periodic = TRUE)
ipMat <- ibs(px, knots = knots, Boundary.knots = c(0, 1),
intercept = TRUE, periodic = TRUE)
dp1Mat <- deriv(pbsMat)
dp2Mat <- deriv(pbsMat, derivs = 2)
par(mfrow = c(1, 2))
plot(pbsMat, ylab = "Periodic B-splines", mark_knots = "boundary")
plot(ipMat, ylab = "The integrals", mark_knots = "boundary")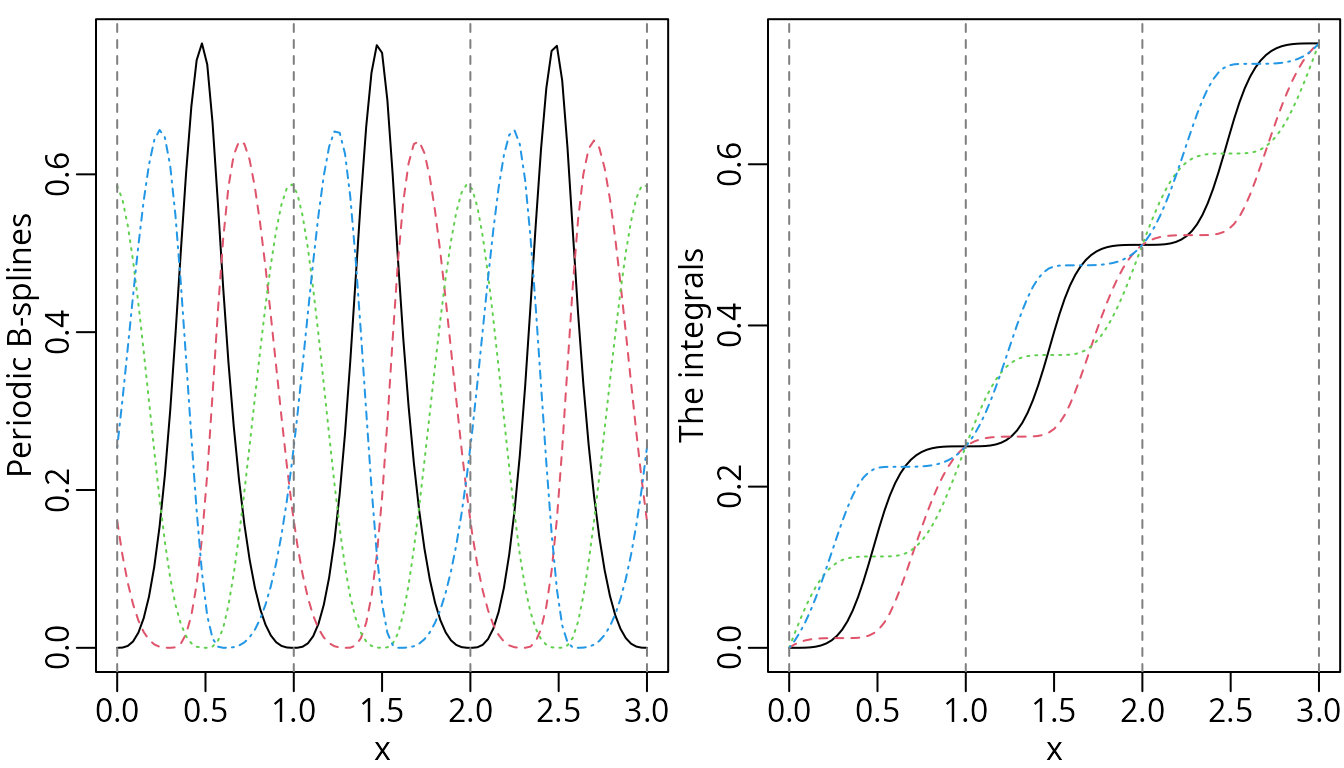
plot(dp1Mat, ylab = "The 1st derivatives", mark_knots = "boundary")
plot(dp2Mat, ylab = "The 2nd derivatives", mark_knots = "boundary")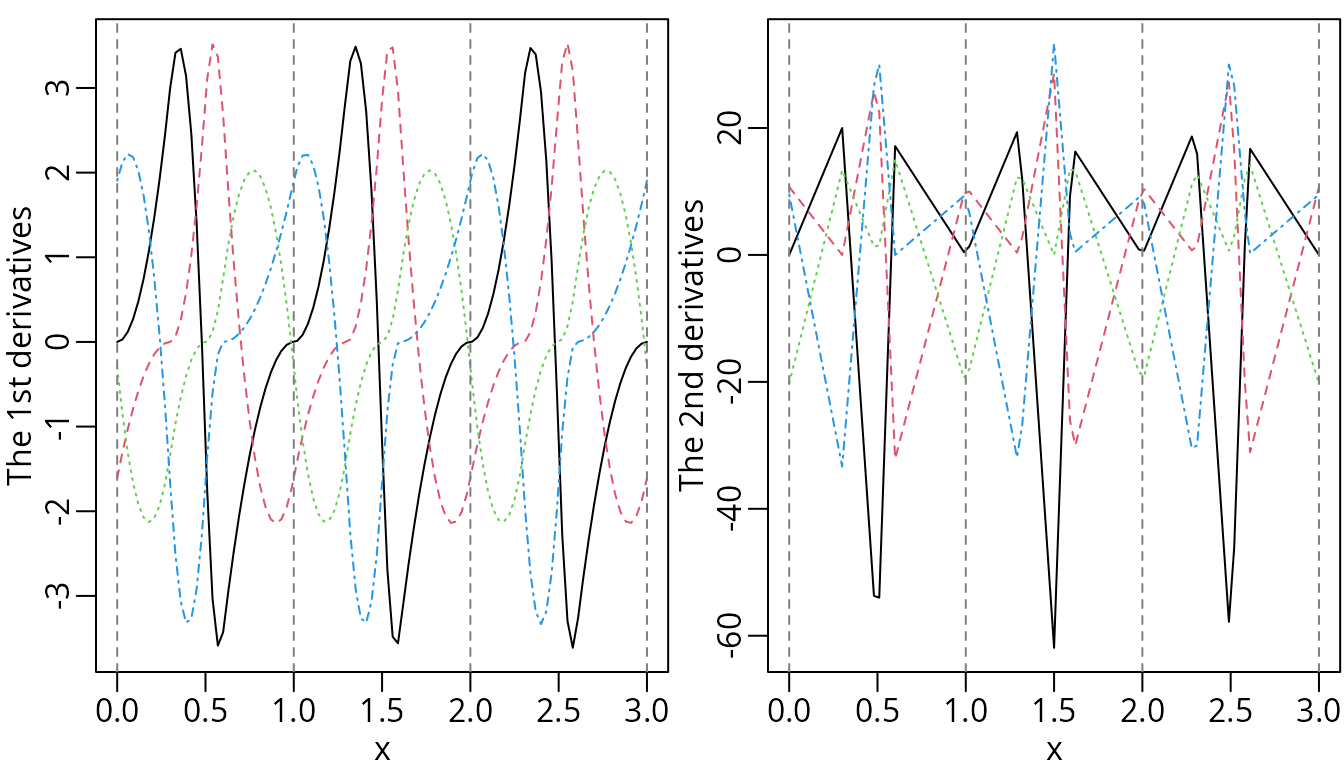
For reference, the corresponding integrals and derivatives are also visualized.
M-Splines
M-spline Basis Functions
M-splines (Ramsay 1988) can be
considered the normalized version of B-splines with unit integral within
boundary knots. An example given by Ramsay
(1988) was a quadratic M-splines with three internal knots placed
at 0.3, 0.5, and 0.6. The default boundary knots are the range of
x, and thus 0 and 1 in this example.
msMat <- mSpline(x, knots = knots, degree = 2, intercept = TRUE)
par(op)
plot(msMat, mark_knots = "all")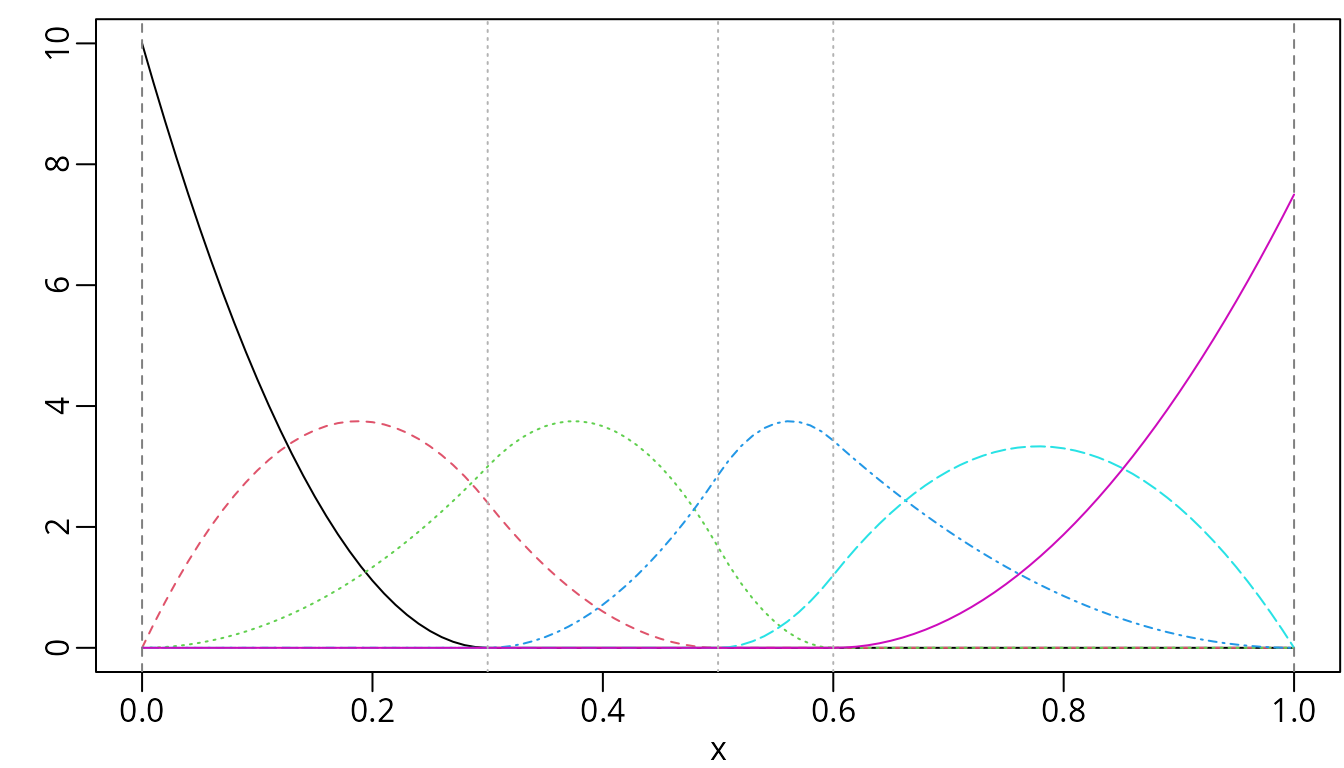
Quadratic M-spline with three internal knots placed at 0.3, 0.5, and 0.6.
The derivative of the given order of M-splines can be obtained by
specifying a positive integer to argument dervis of
mSpline(). Similarly, for an existing mSpline
object generated by mSpline(), one can use the
deriv() method for derivaitives. For example, the first
derivative of the M-splines given in the previous example can be
obtained equivalently as follows:
Periodic M-Splines
The mSpline() function produces periodic splines based
on M-spline basis functions when periodic = TRUE is
specified. The Boundary.knots defines the cyclic interval,
which is the same with the periodic B-splines.
pmsMat <- mSpline(px, knots = knots, intercept = TRUE,
periodic = TRUE, Boundary.knots = c(0, 1))
plot(pmsMat, ylab = "Periodic Basis", mark_knots = "boundary")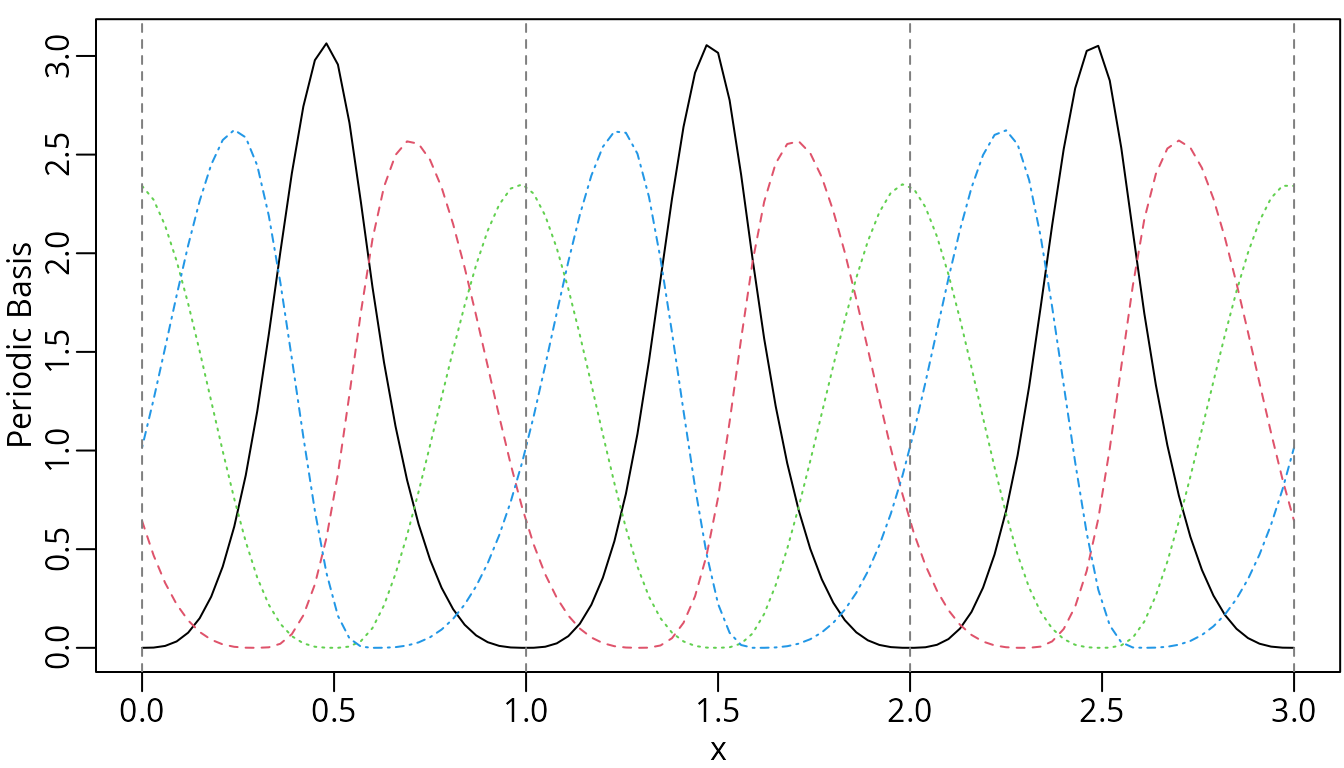
Cubic periodic M-splines.
We may still specify the argument derivs in
mSpline() or use the corresponding deriv()
method to obtain the derivatives when periodic = TRUE.
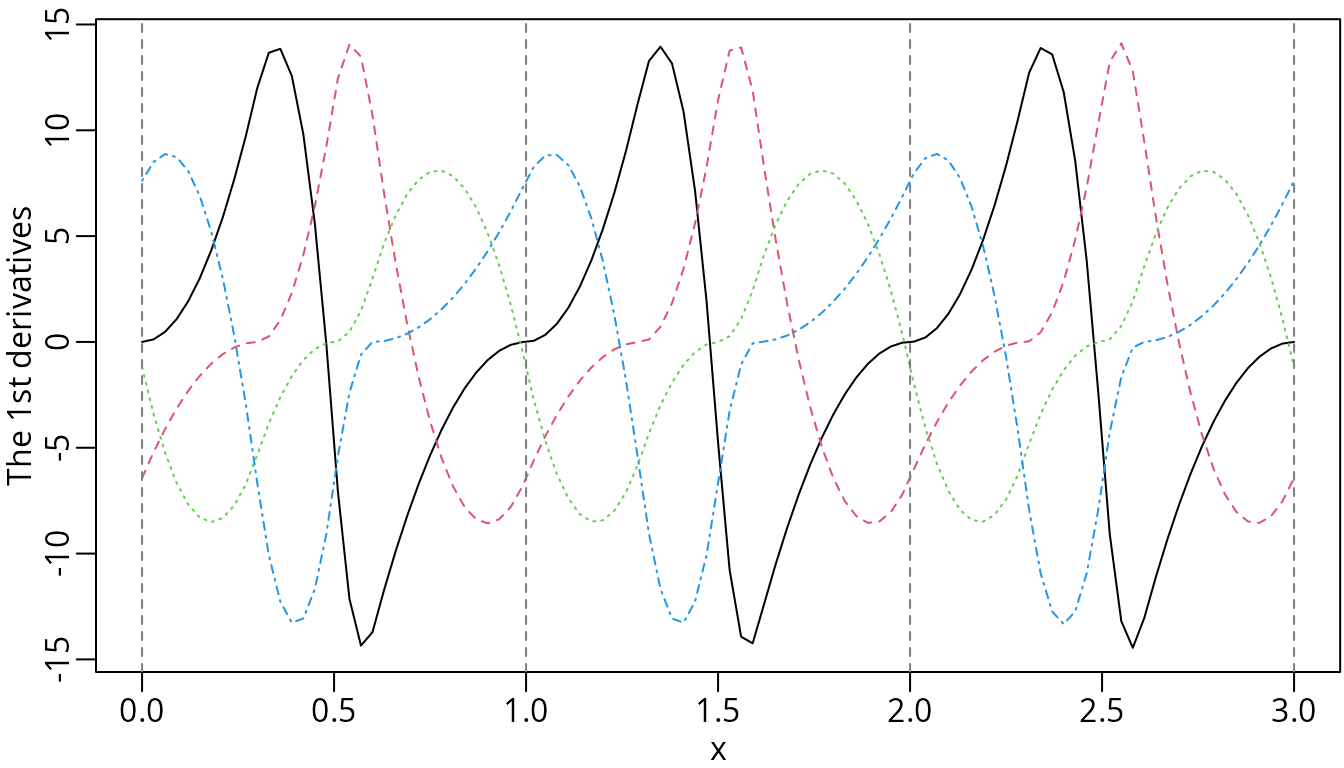
The first derivatives of the periodic M-splines.
Furthermore, we can obtain the integrals of the periodic M-splines by
specifying integral = TRUE. The integral is integrated from
the left boundary knot.
ipmsMat <- mSpline(px, knots = knots, intercept = TRUE,
periodic = TRUE, Boundary.knots = c(0, 1), integral = TRUE)
plot(ipmsMat, ylab = "Integrals", mark_knots = "boundary")
abline(h = seq.int(0, 3), lty = 2, col = "gray")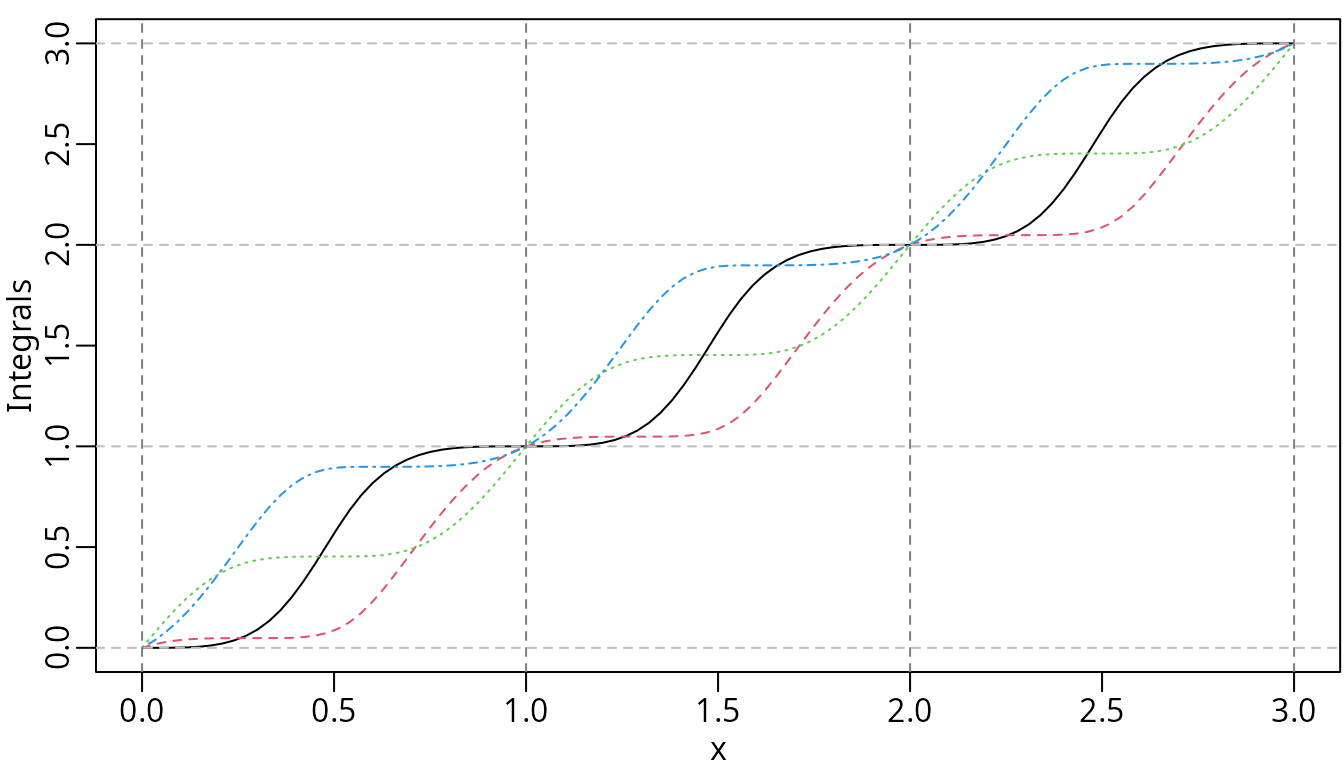
The integrals of the periodic M-splines.
I-Splines
I-splines (Ramsay 1988) are simply the integral of M-splines and thus monotonically nondecreasing with unit maximum value. A monotonically nondecreasing (nonincreasing) function can be fitted by a linear combination of I-spline basis functions with nonnegative (nonpositive) coefficients plus a constant, where the coefficient of the constant is unconstrained.
The example given by Ramsay (1988) was the I-splines corresponding to the quadratic M-splines with three internal knots placed at 0.3, 0.5, and 0.6. Notice that the degree of I-splines is defined from the associated M-splines instead of their polynomial degree.
isMat <- iSpline(x, knots = knots, degree = 2, intercept = TRUE)
plot(isMat, mark_knots = "internal")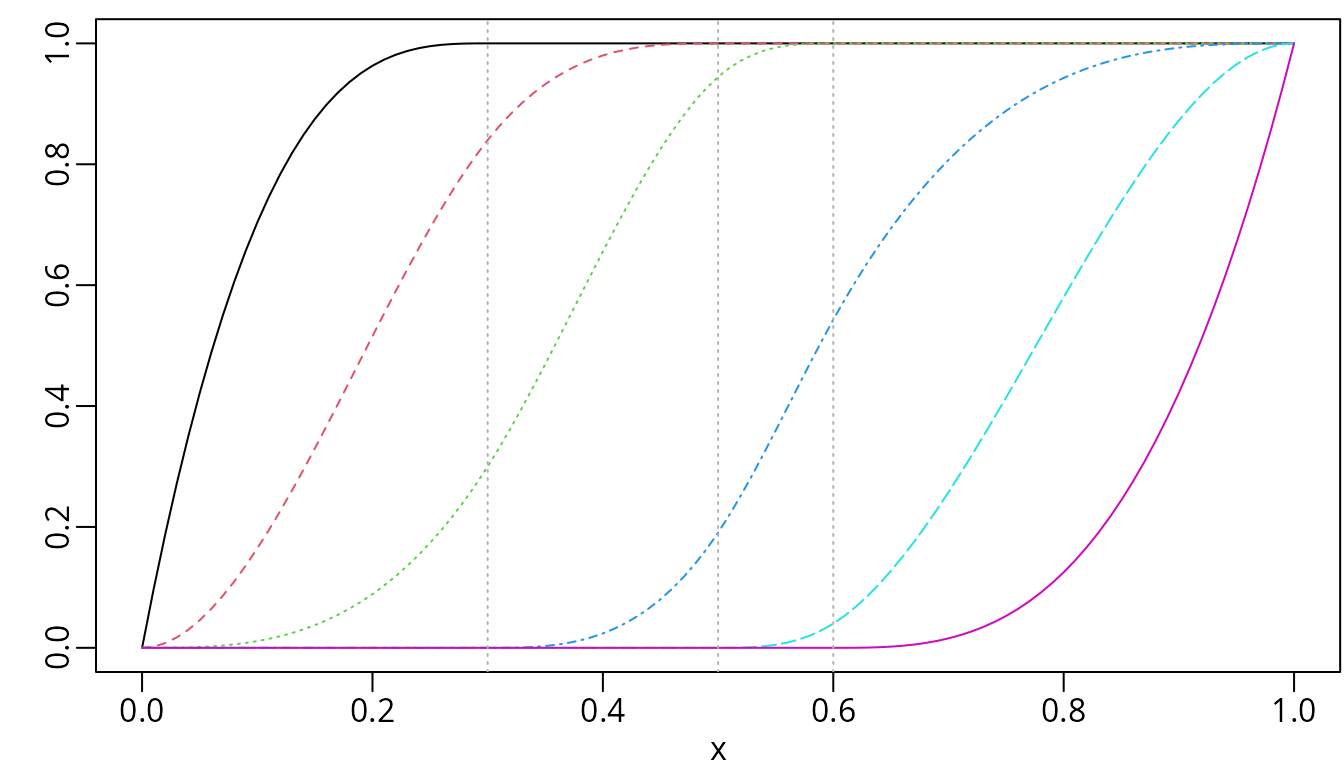
I-splines of degree two with three internal knots placed at 0.3, 0.5, and 0.6.
The corresponding M-spline basis matrix can be obtained easily as the
first derivatives of the I-splines by the deriv()
method.
We may specify derivs = 2 in the deriv()
method for the second derivatives of the I-splines, which are equivalent
to the first derivatives of the corresponding M-splines.
C-Splines
Convex splines (Meyer 2008) called C-splines are scaled integrals of I-splines with unit maximum value at the right boundary knot. Meyer (2008) applied C-splines to shape-restricted regression analysis. The monotone (nondecreasing) property of I-spines ensures the convexity of C-splines. A convex regression function can be estimated using linear combinations of the C-spline basis functions with nonnegative coefficients, plus an unconstrained linear combination of a constant and an identity function . If the underlying regression function is both increasing and convex, the coefficient on the identity function is restricted to be nonnegative as well.
We may specify the argument scale = FALSE in the
function cSpline() to disable the scaling of the integrals
of I-splines. Then the actual integrals of the corresponding I-splines
will be returned. If scale = TRUE (by default), each
C-spline basis is scaled to have unit height at the right boundary
knot.
csMat1 <- cSpline(x, knots = knots, degree = 2, intercept = TRUE)
plot(csMat1)
abline(h = 1, v = knots, lty = 2, col = "gray")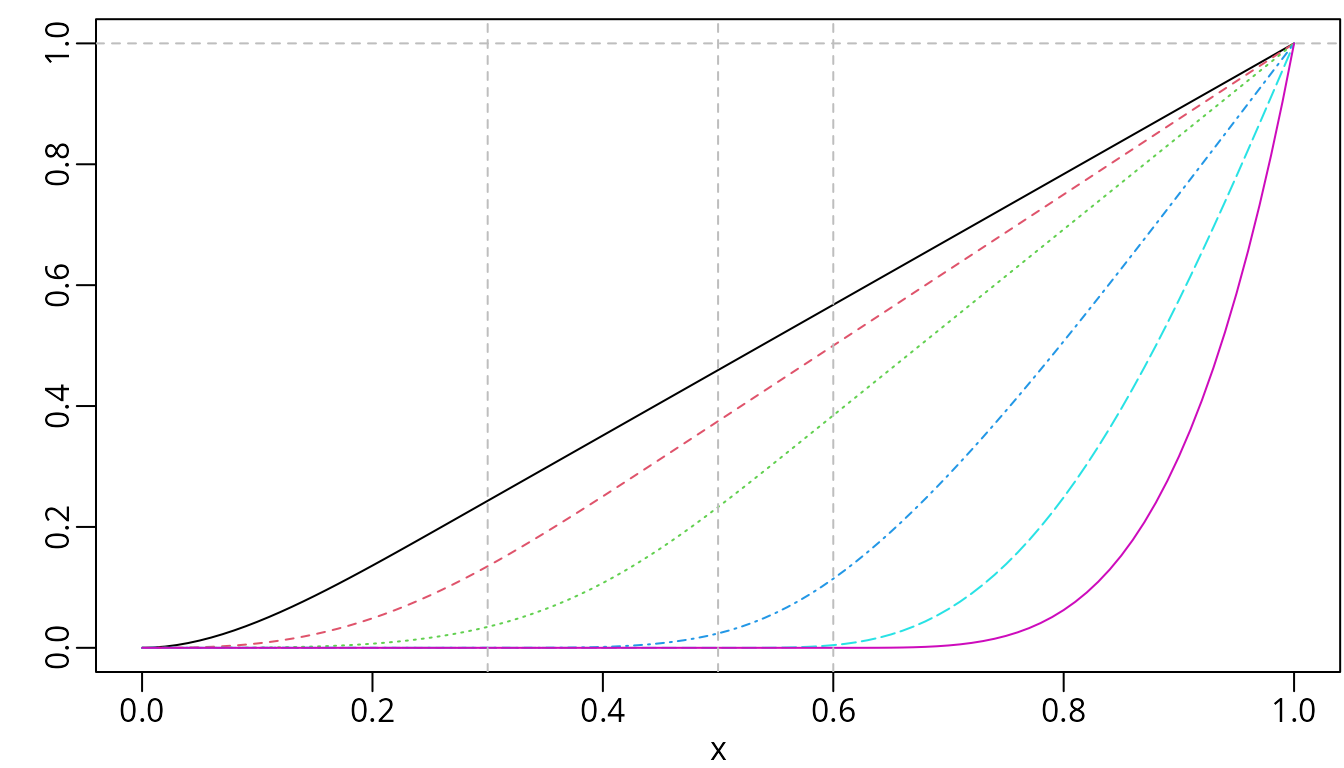
C-splines of degree two with three internal knots placed at 0.3, 0.5, and 0.6.
Similarly, the deriv() method can be used to obtain the
derivatives. A nested call of deriv() is supported for
derivatives of a higher order. However, the argument derivs
of the deriv() method can be specified directly for better
computational performance. For example, the first and second derivatives
can be obtained by the following equivalent approaches,
respectively.
Generalized Bernstein Polynomials
The Bernstein polynomials are equivalent to B-splines without internal knots and have also been applied to shape-constrained regression analysis (e.g., Wang and Ghosh 2012). The -th basis of the generalized Bernstein polynomials of degree over is defined as follows: where . It reduces to regular Bernstein polynomials defined over when and .
We may obtain the basis matrix of the generalized using the function
bernsteinPoly(). For example, the Bernstein polynomials of
degree 4 over
and is generated as follows:
x1 <- seq.int(0, 1, 0.01)
x2 <- seq.int(- 1, 1, 0.01)
bpMat1 <- bernsteinPoly(x1, degree = 4, intercept = TRUE)
bpMat2 <- bernsteinPoly(x2, degree = 4, intercept = TRUE)
par(mfrow = c(1, 2))
plot(bpMat1)
plot(bpMat2)![Bernstein polynomials of degree 4 over [0, 1] (left) and the generalized version over [- 1, 1] (right).](splines2-intro_files/figure-html/bp-1-1.png)
Bernstein polynomials of degree 4 over [0, 1] (left) and the generalized version over [- 1, 1] (right).
In addition, we may specify integral = TRUE or
derivs = 1 in bernsteinPoly() for their
integrals or first derivatives, respectively.
ibpMat1 <- bernsteinPoly(x1, degree = 4, intercept = TRUE, integral = TRUE)
ibpMat2 <- bernsteinPoly(x2, degree = 4, intercept = TRUE, integral = TRUE)
dbpMat1 <- bernsteinPoly(x1, degree = 4, intercept = TRUE, derivs = 1)
dbpMat2 <- bernsteinPoly(x2, degree = 4, intercept = TRUE, derivs = 1)
par(mfrow = c(2, 2))
plot(ibpMat1, ylab = "Integrals")
plot(ibpMat2, ylab = "Integrals")
plot(dbpMat1, ylab = "Derivatives")
plot(dbpMat2, ylab = "Derivatives")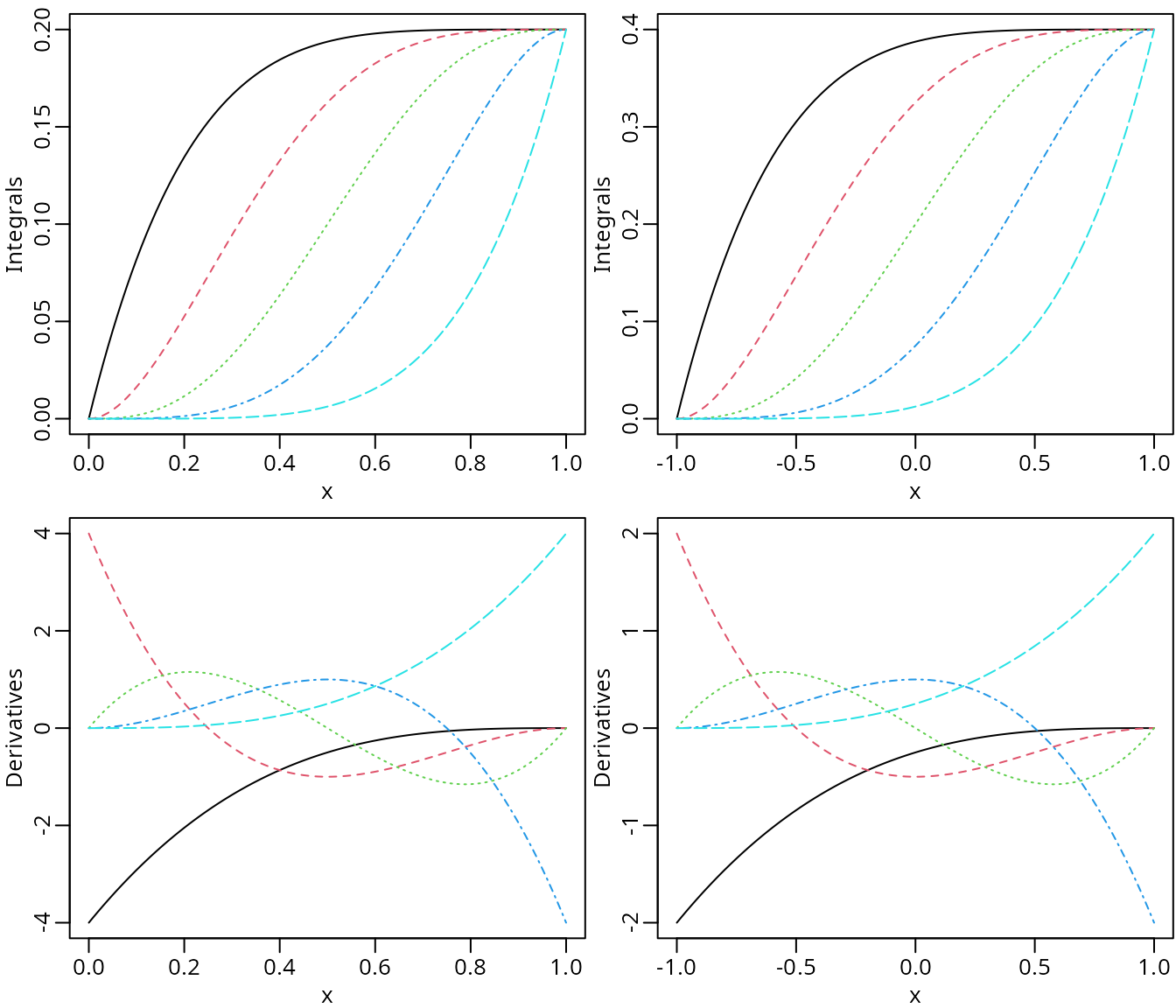
The integrals (upper panel) and the first derivatives (lower panel) of Bernstein polynomials of degree 4.
Similarly, we may also use the deriv() method to get
derivatives of an existing bernsteinPoly object.
stopifnot(is_equivalent(dbpMat1, deriv(bpMat1)))
stopifnot(is_equivalent(dbpMat2, deriv(bpMat2)))
stopifnot(is_equivalent(dbpMat1, deriv(ibpMat1, 2)))
stopifnot(is_equivalent(dbpMat2, deriv(ibpMat2, 2)))Natural Cubic Splines
Nonnegative Natural Cubic Basis Functions
The package provides two variants of the natural cubic splines that
can be constructed by naturalSpline() and
nsk(), respectively, both of which are different from
splines::ns().
The naturalSpline() function returns nonnegative basis
functions (within the boundary) for natural cubic splines by utilizing a
closed-form null space derived from the second derivatives of cubic
B-splines. When integral = TRUE, the function
naturalSpline() returns the integral of each natural spline
basis.
nsMat <- naturalSpline(x, knots = knots, intercept = TRUE)
insMat <- naturalSpline(x, knots = knots, intercept = TRUE, integral = TRUE)
par(mfrow = c(1, 2))
plot(nsMat, ylab = "Basis")
plot(insMat, ylab = "Integrals")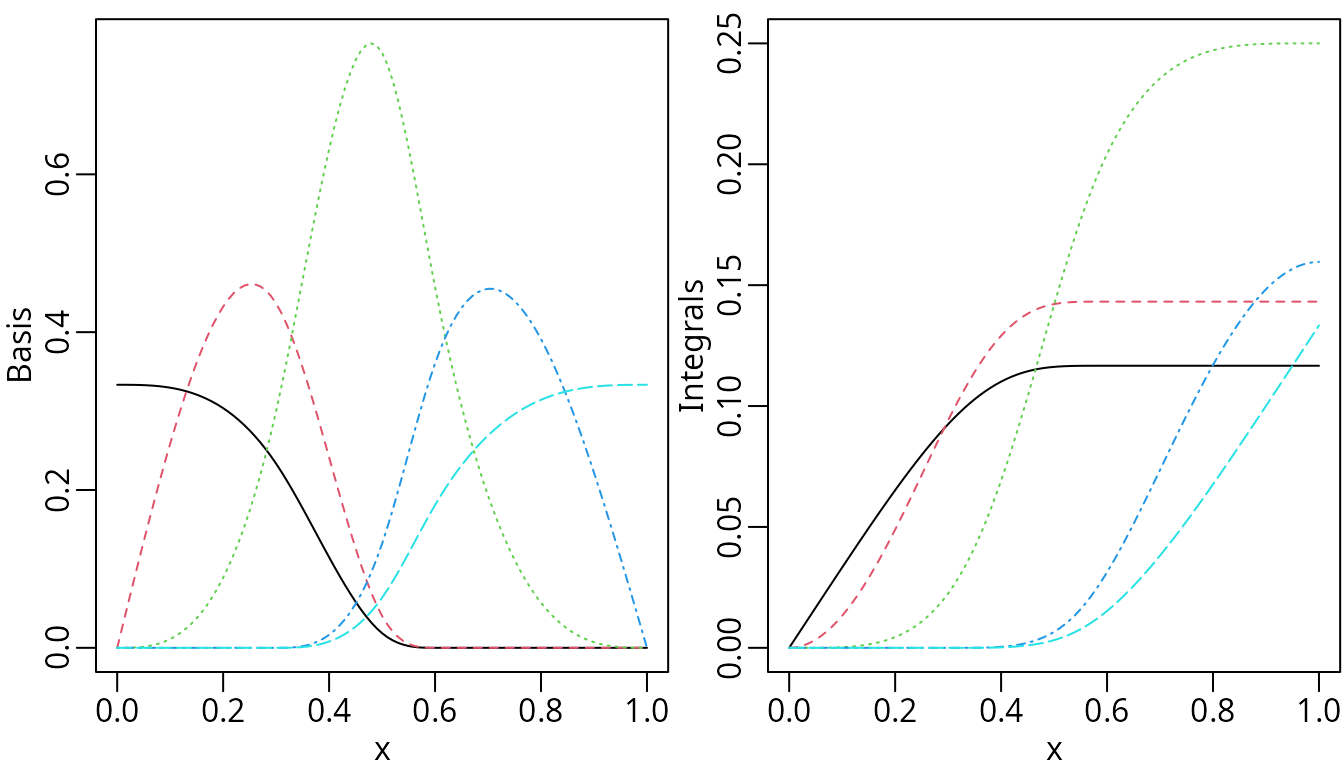
Nonnegative natural cubic splines (left) and corresponding integrals (right).
Similarly, one may directly specify the argument derivs
in naturalSpline() or use the corresponding
deriv() method to obtain the derivatives of spline basis
functions.
d1nsMat <- naturalSpline(x, knots = knots, intercept = TRUE, derivs = 1)
d2nsMat <- deriv(nsMat, 2)
matplot(x, d1nsMat, type = "l", ylab = "The 1st derivatives")
matplot(x, d2nsMat, type = "l", ylab = "The 2nd derivatives")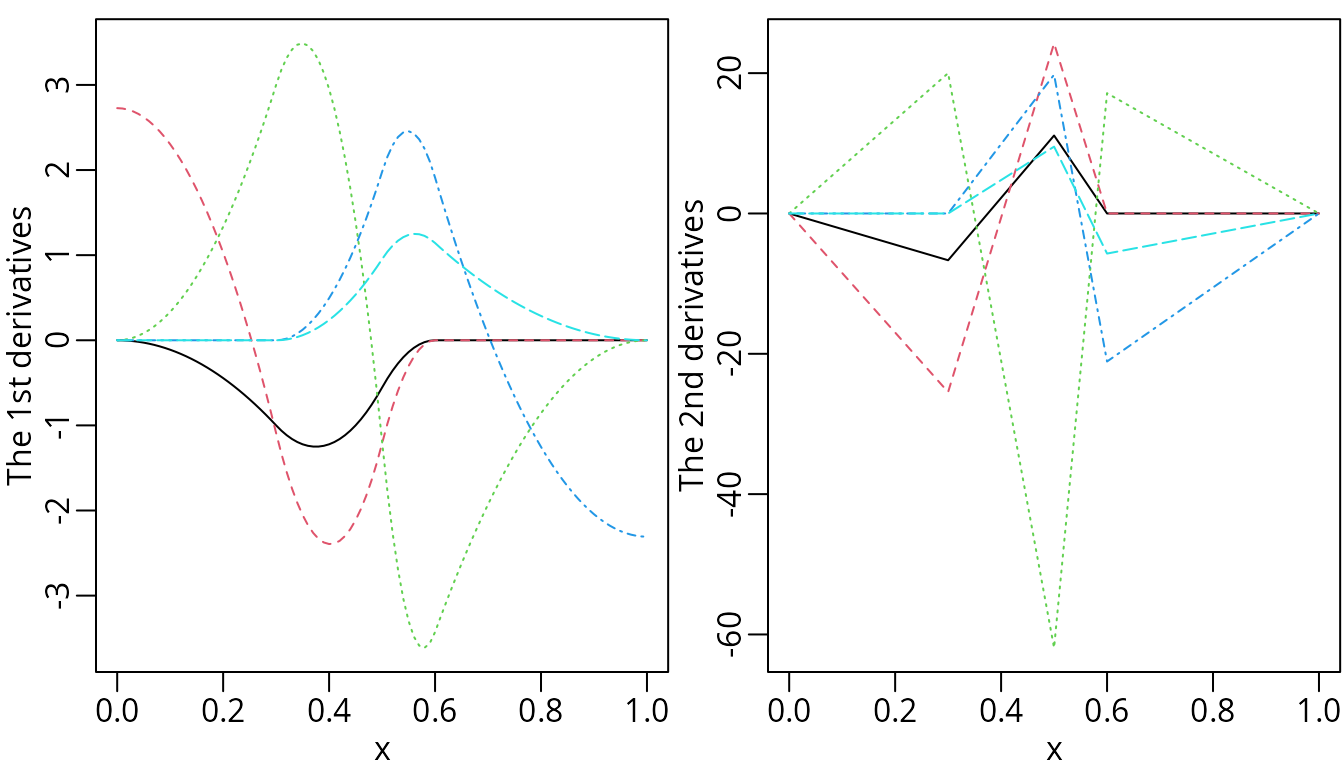
The derivatives of natural cubic splines.
Natural Cubic Basis Functions with Unit Heights at Knots
The function nsk() produces another variant of natural
cubic splines, where only one of the spline basis functions is nonzero
with unit height at every boundary and internal knot. As a result, the
coefficients of the basis functions are the values of the spline
function at the knots, which makes it more straightforward to interpret
the coefficient estimates. This idea originated from the function
nsk() of the survival package (introduced
in version 3.2-8). The implementation of the nsk() of
splines2 essentially follows the
survival::nsk() function. One noticeable argument for
nsk() is trim (equivalent to the argument
b of survival::nsk()). One may specify
trim = 0.05 to exclude 5% of the data from both sides when
setting the boundary knots, which can be a more sensible choice in
practice due to possible outliers. The trim argument is
also available for naturalSpline(), which however is zero
by default for backward compatibility. An illustration of the basis
functions generated by nsk() is as follows:
nskMat <- nsk(x, knots = knots, intercept = TRUE)
par(op)
plot(nskMat, ylab = "nsk()", mark_knots = "all")
abline(h = 1, col = "red", lty = 3)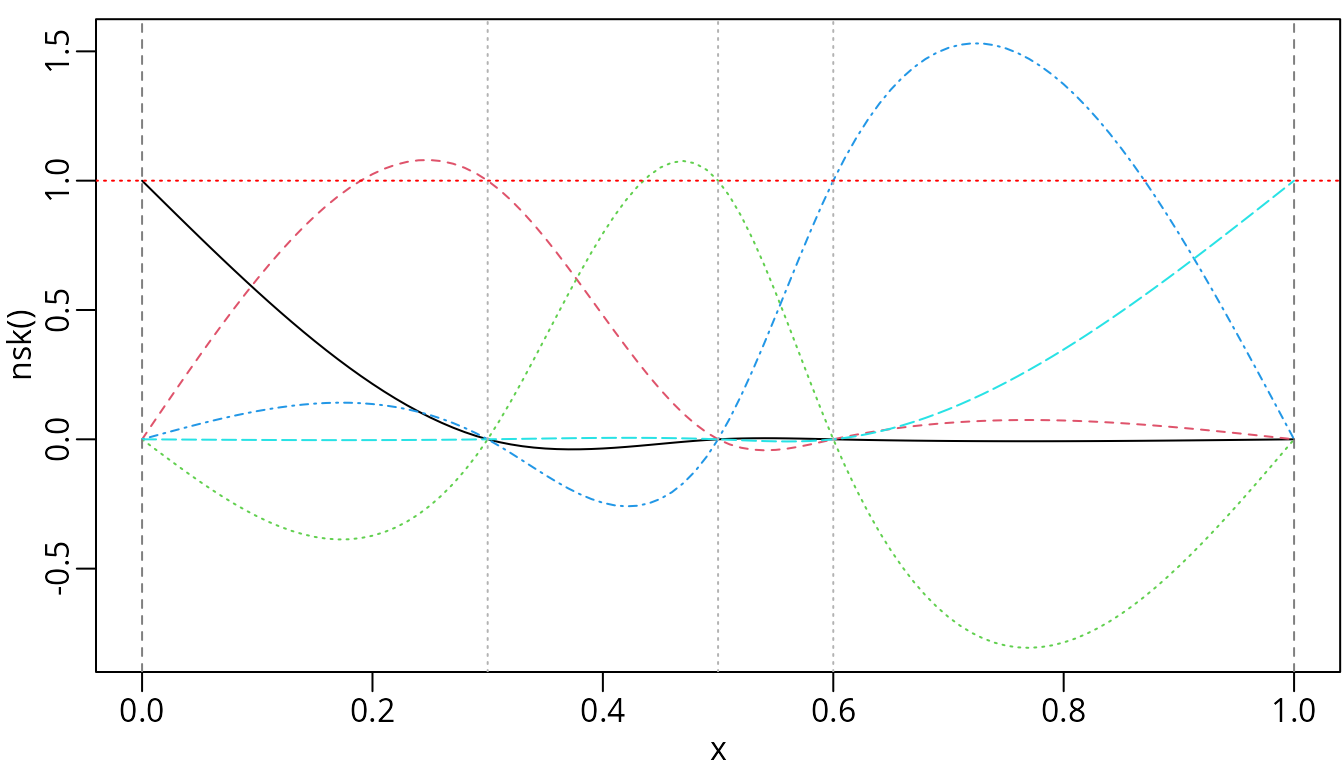
We can visually verify that only one basis function takes a value of one at each knot.
Helper and Alias Functions
Update Spline’s Specification by update()
The update() function is an S3 method to generate spline
basis functions with new x, degree,
knots, or df specifications. The first
argument is an existing splines2 object and additional
named arguments will be passed to the corresponding functions to update
the spline basis functions. Suppose we want to add two more knots to
nskMat for natural cubic spline basis functions and exclude
5% of the data from both sides in total when placing the boundary knots.
We can utilize the update() method as follows:
## [1] 0.2 0.3 0.4 0.5 0.6Evaluation by predict()
The predict() method for splines2 objects
allows one to evaluate the spline function if a coefficient vector is
specified via the coef argument. In addition, it internally
calls the update() method to update the basis functions
before computing the spline function, which can be useful to get the
derivatives of the spline function. If the coef argument is
not specified, the predict() method will be equivalent to
the update() method. For instance, we can compute the first
derivative of the I-spline function from the previous example with a
coefficient vector
seq(0.1, by = 0.1, length.out = ncol(isMat)) at
as follows:
new_x <- c(0.275, 0.525, 0.8)
names(new_x) <- paste0("x=", new_x)
(isMat2 <- predict(isMat, newx = new_x)) # the basis functions at new x## 1 2 3 4 5 6
## x=0.275 0.9994213 0.7730556 0.2310764 0.0000000 0.000000 0.000
## x=0.525 1.0000000 1.0000000 0.9765625 0.2696429 0.000625 0.000
## x=0.8 1.0000000 1.0000000 1.0000000 0.9428571 0.580000 0.125
stopifnot(all.equal(predict(isMat, newx = new_x), update(isMat, x = new_x)))
## compute the first derivative of the I-spline function in different ways
beta <- seq(0.1, by = 0.1, length.out = ncol(isMat))
deriv_ispline1 <- predict(isMat, newx = new_x, coef = beta, derivs = 1)
deriv_ispline2 <- predict(update(isMat, x = new_x, derivs = 1), coef = beta)
deriv_ispline3 <- c(predict(deriv(isMat), newx = new_x) %*% beta)
stopifnot(all.equal(deriv_ispline1, deriv_ispline2))
stopifnot(all.equal(deriv_ispline2, deriv_ispline3))Visualization by plot()
As one may notice in the previous examples, we may visualize the
spline basis functions easily with the plot() method. By
default, the spline basis functions are visualized at 101 equidistant
grid points within the range of x, which can be tweaked by
arguments from, to, and n. In
addition, we can indicate the placement of knots by vertical lines
through the argument mark_knots. The available options are
"none", "internal", "boundary",
and "all". A fitted spline function can be visualized by
specifying the argument coef. An example of
nsk() is as follows:
beta <- seq.int(0.2, length.out = ncol(nskMat), by = 0.2)
plot(nskMat, ylab = "nsk()", mark_knots = "all", coef = beta)
abline(h = beta, col = seq_along(beta), lty = 3)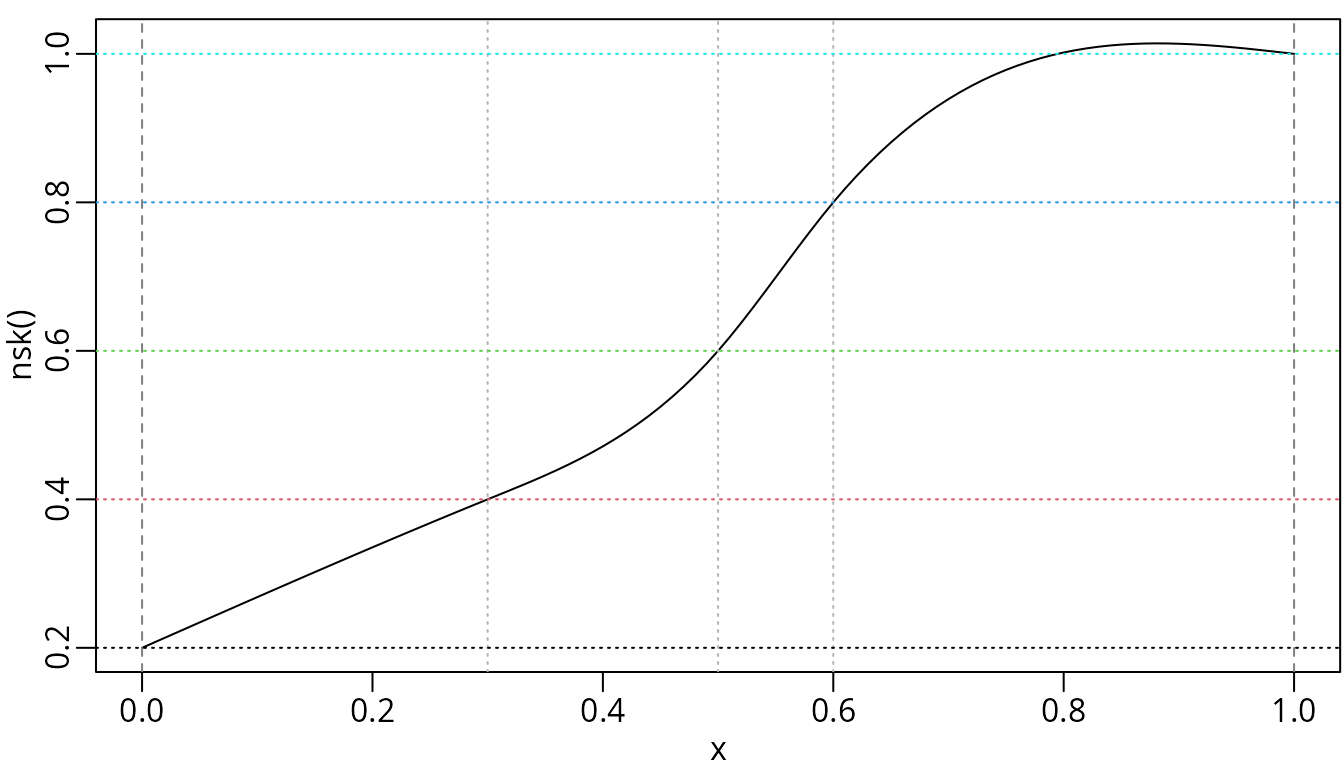
Including Spline Basis Functions in Model Formulas
It is common to directly include spline basis functions in a model formula. To avoid a lengthy model formula, the package provides alias functions that are summarized in the following table:
| Function | Equivalent Alias |
|---|---|
bSpline() |
bsp() |
mSpline() |
msp() |
iSpline() |
isp() |
cSpline() |
csp() |
bernsteinPoly() |
bpoly() |
naturalSpline() |
nsp() |
One may create new alias functions. For example, we can create a new
alias function simply named b() for B-splines and obtain
equivalent models as follows:
b <- bSpline # create an alias for B-splines
mod1 <- lm(weight ~ b(height, degree = 1, df = 3), data = women)
iknots <- with(women, knots(bSpline(height, degree = 1, df = 3)))
mod2 <- lm(weight ~ bSpline(height, degree = 1, knots = iknots), data = women)
pred1 <- predict(mod1, head(women, 10))
pred2 <- predict(mod2, head(women, 10))
stopifnot(all.equal(pred1, pred2))Nevertheless, there is a possible pitfall when using a customized wrapper function for spline basis functions along with a data-dependent placement of knots. When we make model predictions for a given new data, the placement of the internal/boundary knots can be different from the original placement that depends on the training set. As a result, the spline basis functions generated for prediction may not be the same as the counterparts used in the model fitting. A simple example is as follows:
## generates quadratic spline basis functions based on log(x)
qbs <- function(x, ...) {
splines2::bSpline(log(x), ..., degree = 2)
}
mod3 <- lm(weight ~ qbs(height, df = 5), data = women)
mod4 <- lm(weight ~ bsp(log(height), degree = 2, df = 5), data = women)
stopifnot(all.equal(unname(coef(mod3)), unname(coef(mod4)))) # the same coef
pred3 <- predict(mod3, head(women, 10))
pred4 <- predict(mod4, head(women, 10))
all.equal(pred3, pred4)## [1] "Mean relative difference: 0.07185939"
pred0 <- predict(qbs(women$height, df = 5),
newx = head(log(women$height), 10),
coef = coef(mod3)[- 1]) + coef(mod3)[1]
stopifnot(all.equal(pred0, pred4, check.names = FALSE))Although the coefficient estimates are the same, the prediction
results by using the predict.lm() differ. Using an alias
function in the model formula produces correct results.
To resolve this issue, we can create an S3 method for
stats::makepredictcall() as follows:
## generates quadratic spline basis functions based on log(x) with a new class
qbs <- function(x, ...) {
res <- splines2::bSpline(log(x), ..., degree = 2)
class(res) <- c("qbs", class(res))
return(res)
}
## a utility to help model.frame() create the right matrices
makepredictcall.qbs <- function(var, call) {
if (as.character(call)[1L] == "qbs" ||
(is.call(call) && identical(eval(call[[1L]]), qbs))) {
at <- attributes(var)[c("knots", "Boundary.knots", "intercept",
"periodic", "derivs", "integral")]
call <- call[1L:2L]
call[names(at)] <- at
}
call
}
## the same example
mod3 <- lm(weight ~ qbs(height, df = 5), data = women)
mod4 <- lm(weight ~ bsp(log(height), degree = 2, df = 5), data = women)
stopifnot(all.equal(unname(coef(mod3)), unname(coef(mod4)))) # the same coef
pred3 <- predict(mod3, head(women, 10))
pred4 <- predict(mod4, head(women, 10))
all.equal(pred3, pred4) # should be TRUE this time## [1] TRUEExtract Specifications by $
The basis specifications are saved as attributes of the returned
splines2 objects, which means that we can extract one of the
specifications by attr(). Alternatively, we can treat
splines2 objects as lists and use the corresponding
$ method. For example, it is straightforward to extract the
specified trim of nskMat2 by
attr(nskMat2, "trim") or simply
nskMat2$trim.
## [1] 0.025 0.025